Microbes that live in rice paddies, northern peat bogs and other previously unexpected environments are among the bacteria that can generate highly toxic methylmercury, researchers at Oak Ridge National Laboratory and the Smithsonian Environmental Research Center have learned.
This finding, published in Environmental Science and Technology, explains why deadly methylated mercury is produced in areas where the neurotoxin’s presence has puzzled researchers for decades. Methylmercury — the most dangerous form of mercury — damages the brain and immune system and is especially harmful to developing embryos. Certain bacteria transform inorganic mercury into toxic methylmercury.
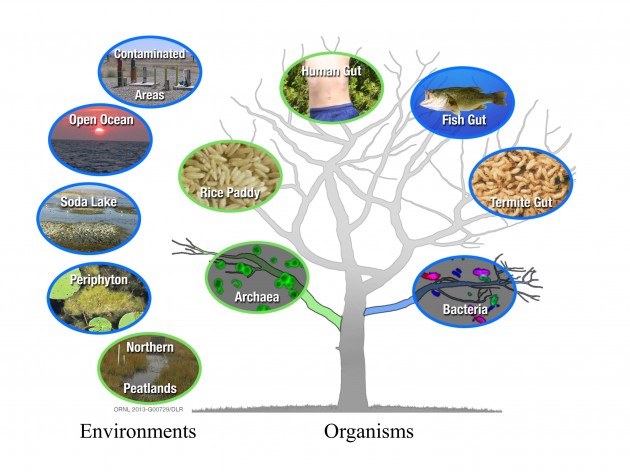
Newly discovered archaeal (green) and bacterial (blue) organisms that are free-living or within another organism on the tree of life, and their larger environments.
The discovery also validates the recent finding that two genes are essential for the methylation of mercury. Previously, only a narrow range of microbes were recognized as mercury methylators, said co-author Dwayne Elias of the Department of Energy laboratory’s Biosciences Division.
“We showed for the first time that many different types of bacteria are able to produce this potent neurotoxin,” Elias said. “The newly identified microbes include methane-producing organisms that live in rice paddies, anaerobic wastewater treatment plants, northern peat lands and possibly within our bodies.”
Elias and colleagues are testing a bacterium from the human intestine that they predict will also methylate mercury. Other bacteria able to transform inorganic into methylmercury include those used in biological dechlorination and metal treatment systems. All of these organisms are anaerobic, which means they grow in habitats without oxygen, including aquatic sediments and wetland soils.
By identifying these organisms, the researchers may have explained why methylmercury is accumulating in unexpected places. The discovery may also help clarify how methylmercury is produced in the open ocean, according to co-author Cindy Gilmour of the Smithsonian Environmental Research Center. Gilmour noted that the newly identified fermentative bacteria may be common in the low oxygen zones of the ocean where methylmercury production occurs. Ocean fish are the primary source of methylmercury in human diets worldwide.
Gilmour and Elias believe this work will have far-reaching implications for understanding the global mercury cycle. Scientists can now use the mercury-methylating organisms and their genes as global biomarkers and develop detection techniques to assess and eventually predict the extent of methylmercury production in a given environment.
“Based on the substantially expanded diversity of organisms and environments for methylation, a number of important habitats deserve further attention,” Gilmour said. “We now have a way to identify and track the abundance of mercury-methylating organisms within these environments, and we hope that the information can be used to devise strategies to minimize methylmercury production.”
Mercury is a global pollutant, released to the atmosphere through coal burning and other industrial uses, and through natural processes. Most of the harm comes from methylmercury bioaccumulation, which is the buildup of the element in tissue that occurs when moving up the food chain.
Other authors of the paper, titled “Mercury methylation by novel microbes from new environments,” are Mircea Podar, Richard Hurt, Steven Brown, Anil Somenahally, Alex Johs and Kathryn Bailey of ORNL, Allyson Bullock of the Smithsonian Environmental Research Center and Andrew Graham of Grinnell College. The paper is available at https://pubs.acs.org/doi/abs/10.1021/es403075t. Funding was provided by DOE’s Office of Science, the National Science Foundation and the Smithsonian Postdoctoral Fellowship Program. Source: Oak Ridge National Laboratory





